Authors
Daniela Vallejo, Jose Luis Castaño, Víctor Arenas, Juan José Domínguez, Lucrecia Yáñez, and Carlos Pipaón.
Introduction
Chronic lymphocytic leukemia (CLL) is the most prevalent form of leukemia in adults. It is characterized by the accumulation of mature B lymphocytes in the blood, bone marrow, and lymphoid tissues. The overexpression of the RRAS2 gene, present in 82% of patients, has been described as the most likely molecular cause of the disease. Although the transformation by RRAS2 has been traditionally associated with genetic mutations, recent studies indicate that its overexpression without mutations is sufficient to induce CLL.
On the other hand, two other molecular mechanisms, i.e. the elevation of the ubiquitin-like post-translational modifications (UBL-PTM) and the increment in RNA translation of certain oncogenes, have demonstrated a high prevalence and relevance in the maintenance of the tumoral phenotype of CLL cells.
Objective
Our objective is to shed light on the role of the aberrant UBL-PTM of RRAS2 found in B-CLL cells on the pathogenesis mediated by RRAS2, in search for new therapeutic targets.
Methods
We have cloned the ORF of RRAS2 and generated mutants of the lysines aberrantly modified in CLL. Preliminary transfection experiments have been performed in HEK293T cells. Transcription of the RRAS2 gene was also analyzed by RT-qPCR and luciferase assays. Then, MEC-1 cell line and ex-vivo experiments on B lymphocytes from patients are utilized to try to extrapolate the data.
Results
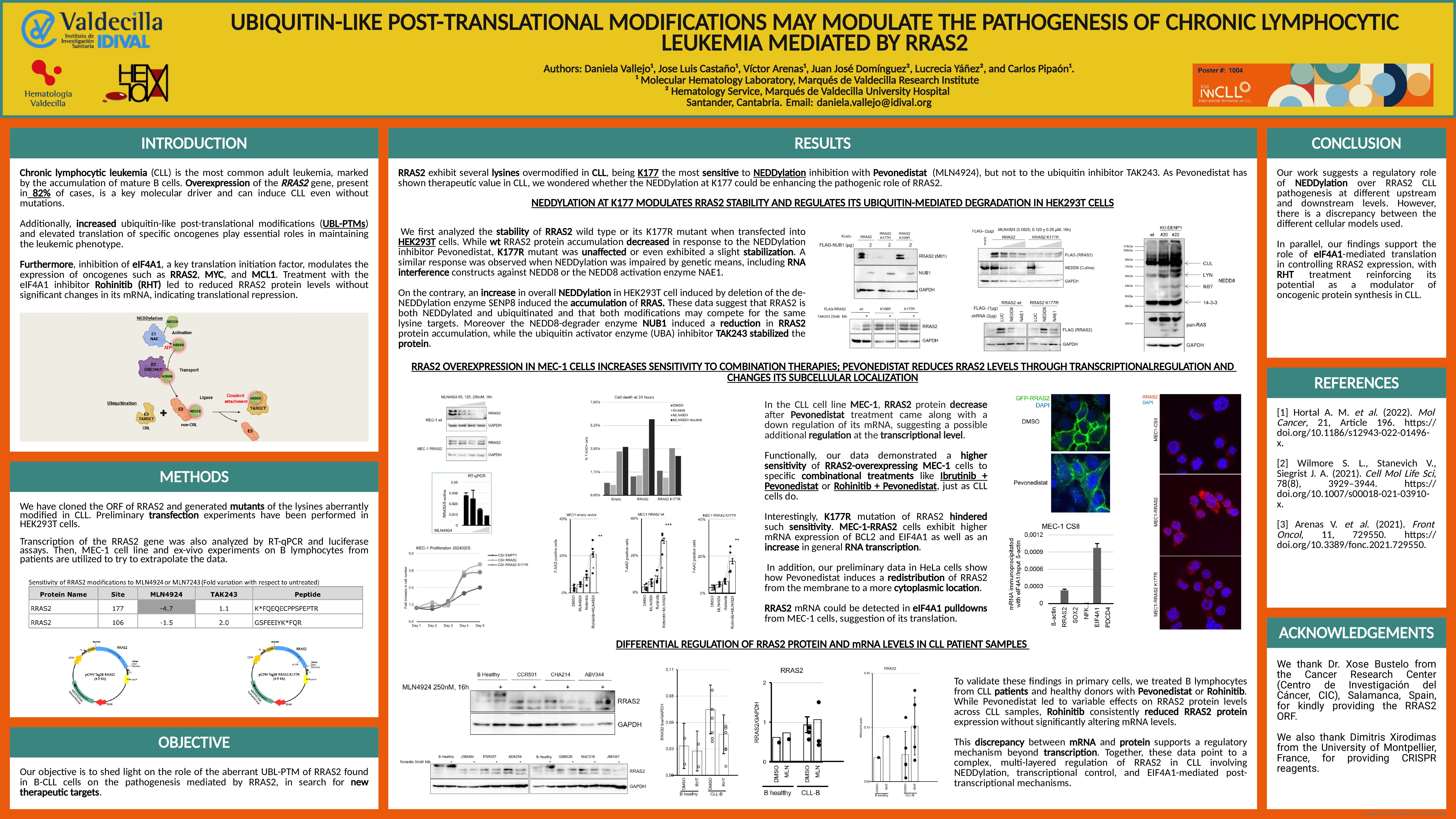
RRAS2 exhibit several lysines overmodified in CLL, being K177 the most sensitive to NEDDylation inhibition with Pevonedistat, but not to the ubiquitin inhibitor TAK243. As Pevonedistat has shown therapeutic value in CLL, we wondered whether the NEDDylation at K177 could be potentiating the pathogenetic potential of RRAS2. We first analyzed the stability of RRAS2 wild type or its K177R mutant when transfected into HEK293T cells. While wt RRAS2 protein accumulation decreased in response to the NEDDylation inhibitor Pevonedistat, K177R mutant was unaffected or even exhibited a slight stabilization. A similar response was observed when NEDDylation was impaired by genetic means, including RNA interference constructs against NEDD8 or the NEDD8 activation enzyme NAE1. On the contrary, an increase in overall NEDDylation in HEK293T cell induced by deletion of the de-NEDDylation enzyme SENP8 induced the accumulation of RRAS2. In concordance with a protective function of NEDDylation against ubiquitin-proteasome degradation, the NEDD8-degrader enzyme NUB1 induced a reduction in RRAS2 protein accumulation, while the ubiquitin activator enzyme (UBA) inhibitor TAK243 stabilized the protein.
In the CLL cell line MEC-1, RRAS2 protein decrease after Pevonedistat treatment came along with a down regulation of its mRNA., suggesting a possible additional regulation at the transcriptional level.
Functionally, our data demonstrated a higher sensitivity of RRAS2-overexpressing MEC-1 cells to specific combinational treatments like Ibrutinib + Pevonedistat or Rohinitib + Pevonedistat, just as CLL cells do. Interestingly, K177R mutation of RRAS2 hindered such sensitivity. MEC-1-RRAS2 cells exhibit higher mRNA expression of BCL2 and EIF4A1 as well as an increase in general RNA transcription. In addition, our preliminary data in HeLa cells show how Pevonedistat induces a redistribution of RRAS2 from the membrane to a more cytoplasmic location.
Conclusion
Our work suggests a regulatory function of NEDDylation over RRAS2 CLL pathogenesis at different upstream and downstream levels.
Keywords : NEDDylation, RNA translation, proteostasis
Please indicate how this research was funded. :
Please indicate the name of the funding organization.: