Authors
Abigail Clark, Joseph Slupsky, Nagesh Kalakonda, Mark Glenn.
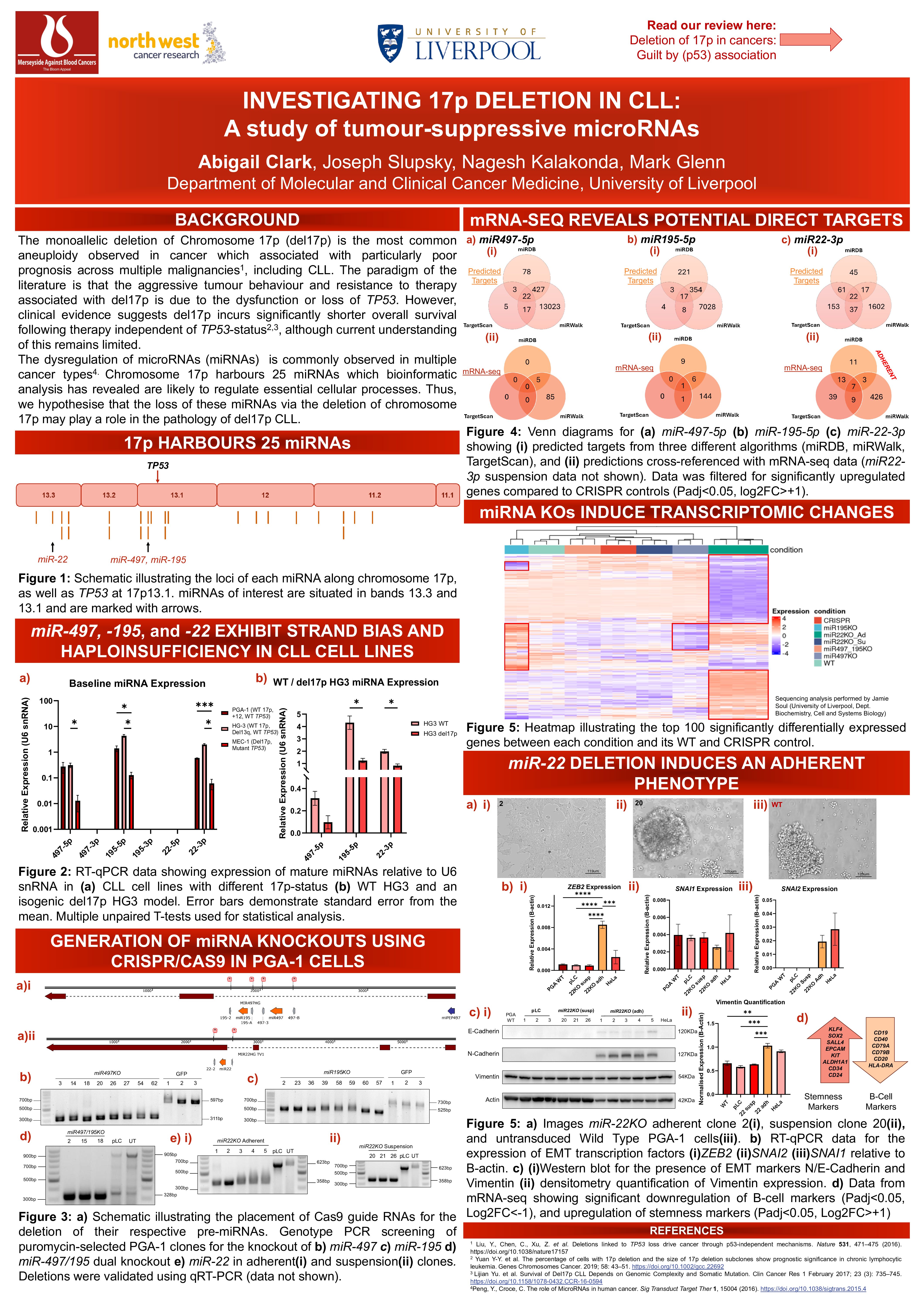
Chronic Lymphocytic Leukaemia (CLL) is a malignancy characterised by the aberrant accumulation of mature-appearing B-cells within lymphoid tissues, resulting in symptoms such as splenomegaly, lymphadenopathy, high white cell counts, and leukaemic infiltration of the bone marrow. It is widely observed that (CLL) is a heterogenous disease which displays strikingly variable clinical courses between patients. Approximately 80% of CLL patients harbour malignant cells with chromosomal aberrations, including the deletions of chromosomes 13q, 11q, and 17p, and trisomy 12q (Döhner et al., 2000), each of which incur differing clinical outcomes. The monoallelic deletion of Chromosome 17p (del17p) is the most common form of aneuploidy observed in both solid and haemic cancers (Chen et al., 2021) and is associated with particularly poor patient prognosis across multiple malignancies, including CLL. In the current literature, these poor outcomes have been mainly attributed to the loss or dysfunction of TP53, which is situated on 17p13.1. This is challenged, however, by clinical studies within CLL demonstrating that del17p results in significantly shorter overall- and treatment-free-survival independent of TP53 status (Brieghel et al., 2019; Yu et al., 2017). Thus, other elements along chromosome 17p may play a role in CLL disease pathology, though current understanding of this remains limited.
Chromosome 17p is particularly gene-rich, and, in addition to ~350 protein-coding genes, there exist multiple non-coding elements including 25 genes encoding microRNAs (miRNAs/miRs), which were of particular focus in this research. miRNAs are small non-coding RNA transcripts ranging from 19-22 nucleotides, which regulate the expression of multiple target genes via mRNA interference, and their dysregulation in cancers is becoming increasingly recognised across multiple malignancies (reviewed in (Reddy, 2015)). To the best of our knowledge, prior to this study, no work has been published regarding the function of 17p-derived miRNAs in the context of B-cell malignancies. The aim of this work is to explore the roles of some of these miRNAs, and how their loss via del17p may contribute to poor patient prognosis.
As a result of preliminary bioinformatic and literature searches, the current study focusses on understanding the roles of tumour suppressor miRNAs: miR497, miR195, and miR22 in the context of CLL. Our data show that these miRs display a consistent strand bias across different CLL cell lines, and are expressed in a gene-dosage dependent manner and may therefore exhibit haploinsufficiency in a del17p karyotype. Using the CLL cell line, PGA-1, we have successfully generated isogenic cell lines using CRISPR/Cas9 technology to knock out each miRNA individually, as well as creating a dual knockout of miR497 and miR195. The knockout of both the individual and dual miR497/miR195 incurred no observable change in phenotype. However, the knockout of miR22 resulted in a distinctive shift to an adherent phenotype, a phenomenon not previously reported for CLL cells. Further experimental investigation revealed a small shift in cell cycle progression in both the individual and dual miR497/miR195, while the adherent miR22 knockouts demonstrated a dramatic drop in the proportion of cells in S-phase, and an increase of cells in G2 phase compared to CRISPR controls.
Preliminary analysis of RNA-sequencing performed on all generated isogenic clones highlighted potential direct targets of each miRNA in conjunction with prediction data. Interestingly, the adherent miR22 knockout cells displayed the most overlap between sequencing and prediction data. The exploration of differentially expressed genes between the dual and individual miR497/miR195 knockouts has also suggested potential compensatory effects and interplay between these miRNAs. The sequencing data arising from the adherent miR22 knockout lines reflected the drastic phenotypic shift: transcription factors and surface proteins typically associated with Epithelial-Mesenchymal Transition (EMT) demonstrated significant upregulation, which reflected our experimental results. Furthermore, B-cell surface markers such as CD19, CD20, and CD79a/b are significantly downregulated, while stemness markers such as KLF4, SOX2, and SALL4 are significantly upregulated, indicating a potential induction of de-differentiation or stem-like features.
This study identifies 17p-derived microRNAs, particularly miR497, miR195, and miR22, as potential contributors to CLL pathogenesis, with miR22 knockout resulting in a unique adherent phenotype, altered cell cycle dynamics, and transcriptional changes suggestive of EMT and stemness. These findings suggest that the poor prognosis associated with del17p in CLL, and potentially other cancers, may be partly mediated by the loss of non-coding RNAs, independent of TP53.
Keywords : microRNA, 17p-deletion, CLL pathogenesis
Please indicate how this research was funded. : This PhD project is jointly funded by Merseyside Against Blood Cancers (UK) and Northwest Cancer Research (UK).
Please indicate the name of the funding organization.: Merseyside Against Blood Cancers (UK), Northwest Cancer Research (UK).