Authors
Nil Albiol, Miguel Arguello-Tomas, Elionor Lynton, Alba Mora, Maria Laura Blanco, Maria Cerdà, JosepMaria Roncero, Eva Gimeno, Lucía Gómez-Pérez, JosepF. Nomdedéu, Esther Moga, Carol Moreno.
Introduction
Measurable residual disease (MRD) correlates with outcome in a variety of hematological malignancies, including chronic lymphocytic leukemia (CLL). The techniques most frequently employed to detect MRD are flow cytometry (FC) and molecular approaches including the detection of Ig rearrangement by high-throughput sequencing (HTS) or real-time quantitative PCR (RQ-PCR). While cell-free DNA (cfDNA) is widely used in lymphomas, in CLL there is little information on whether cfDNA could be more representative of the tumor burden than other methods. Against this background, we aimed to define the role of cfDNA, tentatively compared to other techniques, to assess MRD in patients with CLL.
Methods
This is a non-randomized, prospective, and observational study in which patients with CLL that required therapy (either treatment-naïve or relapsed/refractory) according to iwCLL guidelines.
Peripheral blood samples were prospectively collected before treatment initiation, at the time of response, and every three months thereafter up to two years. Bone marrow samples were collected at the time of response whenever possible.
MRD was assessed by multiparameter FC (by 6-colour, CD19, CD43, CD20, CD81, CD79b, and CD5) and clonoSEQ®. For the detection of the rearranged IGHV sequence, genomic DNA was extracted from PMN cells and sent to Adaptive alongside with frozen plasma samples. The cut-off used for MRD positivity by HTS was above 10-6 (LOD) and 10-4 in flow MRD samples.
Results
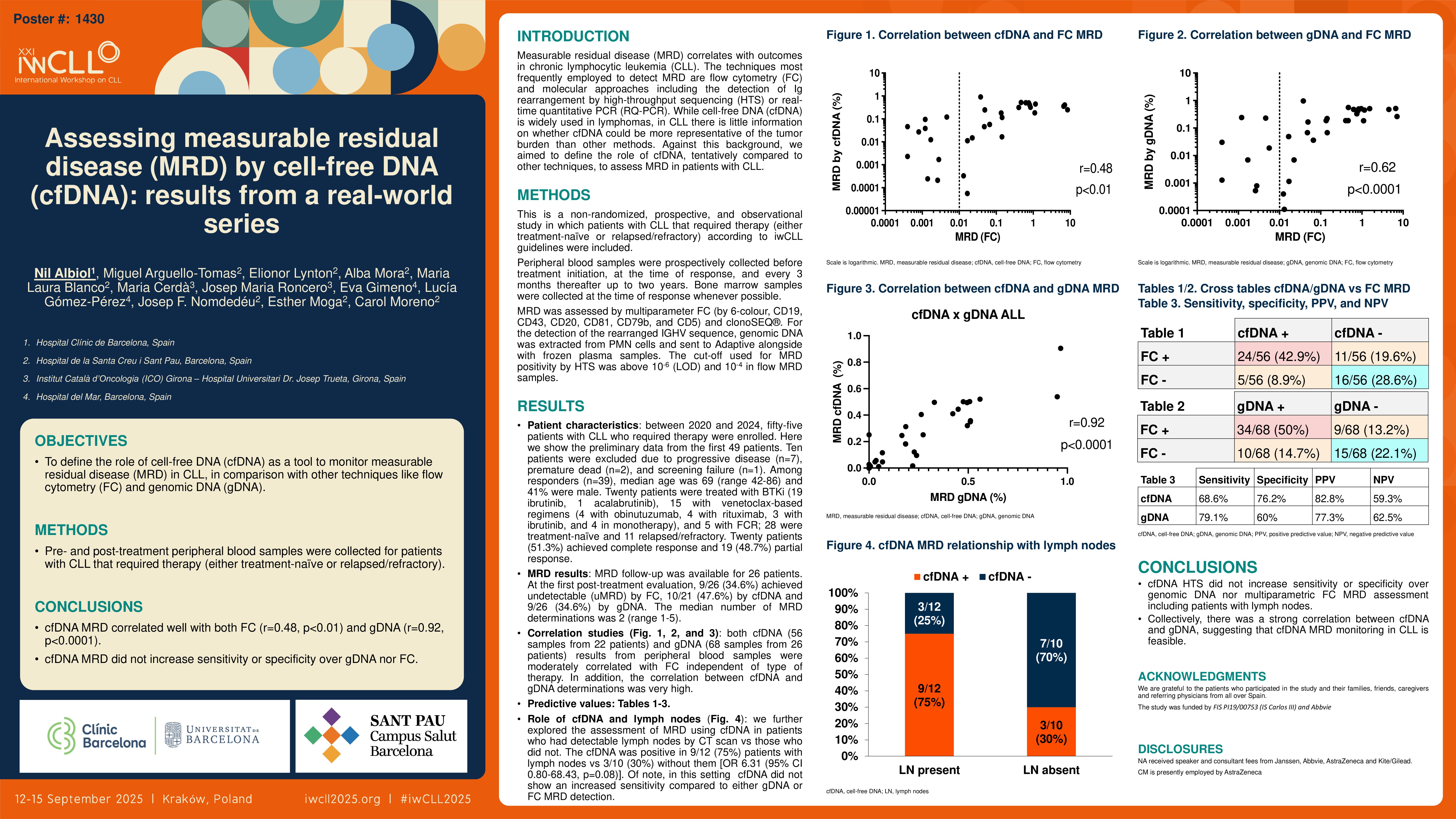
Between 2020 and 2024, fifty-five patients with CLL who required therapy were enrolled. Here we show the preliminary data from the first 49 patients. Ten patients were excluded due to progressive disease (n=7), premature dead (n=2), and screening failure (n=1). Among responders (n=39), median age was 69 (range 42-86) and 41% were male. Twenty patients were treated with BTKi (19 ibrutinib, 1 acalabrutinib), 15 with venetoclax-based regimens (4 with obinutuzumab, 4 with rituximab, 3 with ibrutinib, and 4 in monotherapy), and 5 with FCR; 28 were treatment-naïve and 11 relapsed/refractory. Twenty patients (51.3%) achieved complete response and 19 (48.7%) partial response. At the first post-treatment evaluation, 9/26 (34.6%) achieved undetectable (uMRD) by FC, 10/21 (47.6%) by cfDNA and 9/26 (34.6%) by gDNA. MRD follow-up was available for 26 patients. The median number of MRD determinations was 2 (range 1-5). Both cfDNA (56 samples from 22 patients) and gDNA (68 samples from 26 patients) results from peripheral blood samples were moderately correlated with FC (r=0.48, p< 0.01; r=0.62, p< 0.0001), independent of type of therapy. In addition, the correlation between cfDNA and gDNA determinations was very high (r=0.92, p< 0.0001).
When comparing cfDNA vs FC, the sensitivity and specificity were 68.6% and 76.2%, and for gDNA 79.1% and 60%, respectively. The positive and negative predictive values for cfDNA were 82.8% and 59.3%, and for gDNA they were 77.3% and 62.5%, respectively. There were 16/56 (28.6%) discordant results in cfDNA and 19/68 (28%) in gDNA vs FC.
We further explored the assessment of MRD using cfDNA in patients who had detectable lymph nodes by CT scan vs those who did not. The cfDNA was positive in 9/12 (75%) patients with lymph nodes vs 3/10 (30%) without them [OR 6.31 (95% CI 0.80-68.43, p=0.08)].Of note, in this setting cfDNA did not show an increased sensitivity compared to either gDNA or FC MRD detection.
Conclusions
In summary, these results show that cfDNA HTS did not increase sensitivity or specificity over genomic DNA nor multiparametric FC MRD assessment including patients with lymph nodes. Collectively, there was a strong correlation between cfDNA and gDNA, suggesting that cfDNA MRD monitoring in CLL is feasible.
Keywords : cfDNA, MRD, clonoSEQ
Please indicate how this research was funded. : Grants FIS PI19/00753 (IS Carlos III) and private research funding from Abbvie
Please indicate the name of the funding organization.: Instituto de Salud Carlos III
Abbvie