Authors
Jun Ma, Yang Zhang, Pan Gao, Peipei Li, Zhumei Zhan, Changqing Zhen, Xianghua Wang, Xing Cui, Dongwei Kang, Xiaosheng Fang, and Ya Zhang.
Background
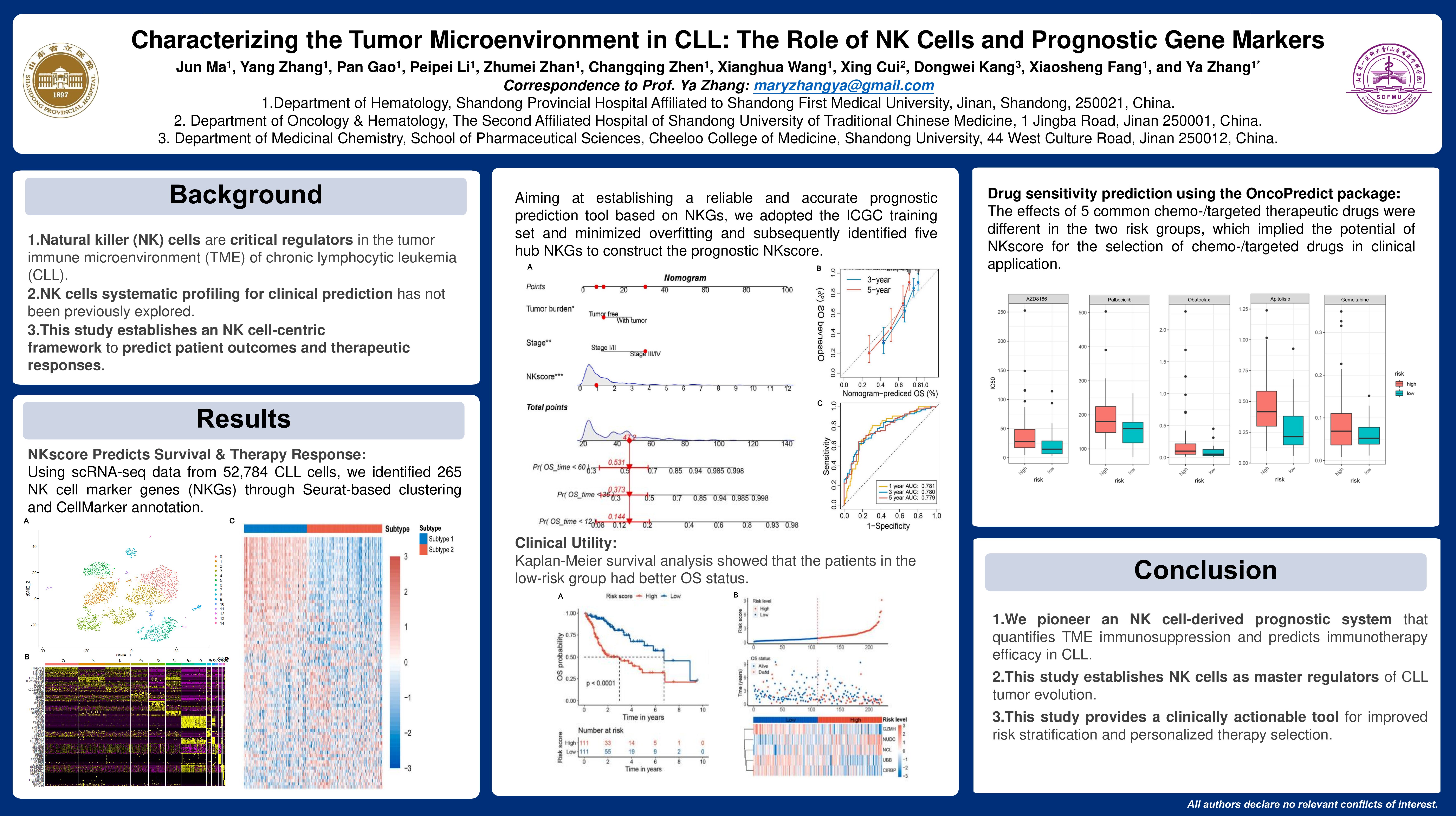
Natural killer (NK) cells are critical regulators within the tumor immune microenvironment (TME) of chronic lymphocytic leukemia (CLL), yet their systematic profiling for clinical prediction remains unexplored. This study establishes an NK cell-centric framework to predict patient outcomes and therapeutic responses.
Methods
Using scRNA-seq data from 52,784 CLL cells, we identified 265 NK cell marker genes (NKGs) through Seurat-based clustering (resolution=0.3) and CellMarker annotation. Prognostic NKGs were refined via multi-split survival analysis (ICGC cohort, n=487) and consensus clustering. Differential expression analysis (|logFC|>1, adj.P.Val < 0.01) and functional enrichment (GO/KEGG) delineated NK subtypes. A five-gene prognostic signature was constructed through 500-replicate LASSO-COX regression and stepwise selection. Immune landscapes were characterized via ESTIMATE scores, TCR repertoire analysis, and immunophenoscore.
Results
Consensus clustering stratified CLL patients into two NK subtypes: Subtype 1 demonstrated superior overall survival (OS, p< 0.001), enriched IGHV mutations (OR=2.1, p=0.007), and heightened immune cytolytic activity, while Subtype 2 exhibited advanced Rai stage predominance (68% vs 32%, p=0.002) and TP53/ATM mutational enrichment (q < 0.01). The five-gene NKscore robustly discriminated high- risk and low-risk groups (median cutoff), with high-risk patients showing significantly worse OS (HR=3.21, 95%CI 2.18-4.73, p< 0.001), progression-free survival (HR=2.67, p< 0.001), and disease-specific survival (HR=4.02, p< 0.001), corroborated by 5-year time-dependent ROC analysis (AUC=0.84). Mechanistic profiling revealed high-NKscore tumors harbored an immunosuppressed microenvironment characterized by depleted TCR clonality (Shannon index: 5.2 vs 8.1, p=0.008), elevated T-cell exhaustion markers, reduced ESTIMATE ImmuneScore (-412 vs +285, p< 0.001), and TGF-β pathway activation (FDR < 0.01), collectively conferring resistance to PD-1/CTLA-4 inhibitors (Immunophenoscore: 0.32 vs 0.68, p=0.004) and ibrutinib (IC50: 1.8μM vs 0.7μM, p=0.02). Conversely, low-NKscore tumors displayed inflamed microenvironments with Th1/Th17 polarization, CXCL9/10 upregulation (>5-fold), and enhanced chemokine signaling. Integration of NKscore into a clinical nomogram significantly improved 5-year survival prediction (C-index: 0.79 vs 0.68, p=0.003).
Conclusion
We pioneer an NK cell-originated prognostic system that quantifies TME immunosuppression and predicts immunotherapy efficacy in CLL. This work positions NK cells as master regulators of tumor evolution and provides a clinically actionable tool for risk stratification and personalized therapy selection.
Keywords : Chronic lymphocytic leukemia; NK cell; Prognosis
Please indicate how this research was funded. : This study was supported by the National Natural Science Foundation (No. 82000195); Shandong Provincial Natural Science Foundation (No. ZR2024MH050, ZR2020QH094); China Postdoctoral Science Foundation (No. 2022M721981); Taishan Scholars Program of Shandong Province (No. tsqn201909184); Shandong Provincial Health Construction Program of Medical Science and Technology Innovation Team; Academic Promotion Programme of Shandong First Medical University (No.2020RC007); Research Promotion Program of Shandong Provincial Hospital.
Please indicate the name of the funding organization.: National Natural Science Foundation of China; Shandong Provincial Natural Science Foundation; China Postdoctoral Science Foundation; Taishan Scholars Program of Shandong Province; Shandong Provincial Health Construction Program of Medical Science and Technology Innovation Team; Shandong First Medical University; Shandong Provincial Hospital.